Merck Finds Commercial Solution in Collaboration
July 25, 2013
By Amy E. Kallmerten and Rudy Potenzone
July 26, 2013 | Commentary | With a growing interest in biologics and a significant investment in deploying an electronic laboratory notebook (ELN) to thousands of employees, Merck decided in 2011 to partner with PerkinElmer to expand the functionality of the informatics provider’s E-Notebook ELN solution to meet the needs of its biologics workflows.
Merck Research Laboratories (MRL) had identified biologics as an area of increasing focus and strategic growth, but noted a gap in the ability of existing enabling technologies and processes to support the structured data capture, analysis, and workflow management that is required for the various complex stages of biologics research and development. Additionally, MRL wanted an integrated solution that would allow users to search, access, and share biologics data and manage tasks. Leveraging its investment in the E-Notebook platform by developing additional functionality to support its biologics R&D was deemed the best path forward.
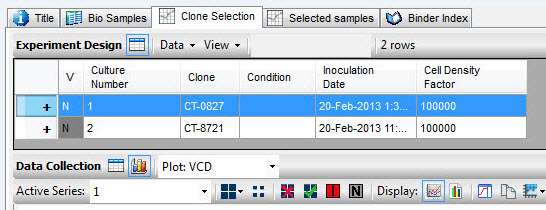 |
| Clone selection experimental design |
“One of the key factors to the success of this process was PerkinElmer’s willingness to collaborate and respond to our needs to enhance a commercial, not custom, product,” said Dermot BarryWalsh, executive director for Vaccine/Biologics R&D at Merck Research Laboratories (MRL). “We did not want substantial custom code that we would need to feed and care for. We wanted a commercial product with support behind it.”
Having merged in 2009 with Schering-Plough, MRL biologics researchers were using a combination of spreadsheets, paper lab notebooks, and the E-Notebook ELN solution from PerkinElmer, which had been introduced as a paper replacement. Merck sought to deploy a standard solution with integrated biologics-specific workflow capabilities, such as attached data structure, taxonomy, or data models. They believed that providing a data structure around research and development candidates would enable results to be searched and shared. The problem was no commercial solution existed. So, after evaluating its options, Merck entered into a collaborative agreement with PerkinElmer to build out the E-Notebook solution as an integrated platform to support MRL’s biologics workflows. The two organizations worked jointly to identify requirements to further develop the E-Notebook BioAssay module with sample tracking and management capabilities for a collaborative biologics environment.
Merck anticipated two levels of benefit from this collaboration:
1. MRL scientists would be able to capture data with content, enabling them to perform sequential and comparative analysis throughout the life cycle of a product.
2. MRL lab managers and others would be able to assess productivity, cycle time, capacity understanding, and perform full product life cycle analysis.
“Because it’s a structured database, we can more easily access and analyze the data than if it’s just attached as an Excel spreadsheet,” said BarryWalsh.
.jpg) |
| Culture profiling experiment design |
To expand functionality to meet MRL’s growing needs, PerkinElmer first evaluated the BioAssay and RDLIMS modules in a “sandbox” environment to access the new functionality and determine if the solutions were fit for purpose. Once the proof of concept was completed, the new functionality was built out and the enterprise solution was tested in a production pilot deployment that began in August 2012. After that pilot was complete, the biologics functionality for the BioAssay and sample tracking and management solutions was rolled out to select users within Merck’s Vaccines Development, Biologics Discovery, and Biologics Development areas. Additionally, researchers in In Vivo data management within Merck’s Discovery and Preclinical R&D groups deployed the E-Notebook BioAssay module for data collection, analysis, and reporting. Because of the broad base of existing E-Notebook users throughout MRL, the enhanced biologics functionality is accessible to various functional groups based on roles and responsibilities via the security system within the ELN.
.jpg) |
| Glucose growth evaluation plot |
Business Justification
MRL’s business justification for the collaborative development project initially focused on productivity gains through the elimination of manual data manipulation, overhead, and lost data or information that occurred in the preexisting environment.
Mark Jankowski, Merck’s director for application services, said the company looks forward to performance enhancements as well from aggregating, analyzing, reviewing, and reporting data and a reduction of low-value experiments due to better visibility of that data. Measuring these improvements first requires a more substantial aggregation of data. At this stage, however, enhancements to workflow management are being realized, according to Merck. A process that would have been completely manual before is now being performed by the E-Notebook’s sample tracking and management functionality, which tracks samples generated by one group and processed by another and then automatically reports results.
Future collaborative plans involve additional, iterative development of the enhanced E-Notebook BioAssay module as new requirements emerge from the pilot and subsequent waves of deployment and use. PerkinElmer and Merck will continue to address the areas of prioritized needs, with particular emphasis placed on enhancing usability and flexibility. Merck says it is further interested in deploying the BioAssay module for its external partners so that outsourced work can be performed within BioAssay and seamlessly rolled back into Merck’s system.
Amy E. Kallmerten is Program Manager, Enterprise R&D and Rudy Potenzone is VP of Product Strategy, Informatics and Business Line Leader, Enterprise R&D for PerkinElmer.


