Ayasdi Seeks Out the Underlying Shapes of Big Data
By Aaron Krol
May 14, 2014 | More than a hundred companies set up booths at the 2014 Bio-IT World Conference & Expo to demonstrate their products, but it seems safe to say only one of them could claim the Miami Heat as a customer.
Ayasdi, whose eclectic client base also includes Merck, General Electric, Citigroup, and the U.S. Department of Agriculture, has actually published a paper in Nature Scientific Reports on the nature of basketball positions, in which the authors propose that the three classic designations of guard, forward, and center can be further divided into thirteen subcategories with distinct and definable styles of play. The company’s flagship product, a data analysis platform, specializes in this kind of splintering, finding meaningful groups within large cohorts where huge numbers of variables may be in play. In the basketball study, Ayasdi considered players’ rebounds, assists, turnovers, steals, blocked shots, personal fouls, and points scored per minute, but in other fields of study, the datasets can sprawl to thousands or even millions of variables.
This is especially true in the life sciences, where researchers and even clinical labs are increasingly looking at patients and tissue samples through the lens of genetics, RNA expression, and proteomics. This yields massive numbers of data points that might be key to understanding why some patients respond better to certain drugs, or have different disease prognoses. Compute power and storage architectures are largely keeping pace with this explosion of information, but analytical tools have been slower to adapt. The search for meaning in biomedical big data tends to be piecemeal, zeroing in on a few of the most promising leads without taking a holistic approach to similarities among the patients themselves.
Pek Lum, Ayasdi’s Vice President of Solutions and Chief Data Scientist, believes her company is ready to change that, and had come to the conference in Boston to promote her company’s growing commitment to biomedicine. For ten years, Lum worked at Stephen Friend’s company Rosetta Inpharmatics developing pattern recognition software for in vitro gene expression data, continuing at Rosetta when the company was acquired by Merck. Since joining Ayasdi in 2010, Lum has helped the company pivot toward drug development and human health.
“We really want to make a difference to healthcare,” she says. “Healthcare, and big pharma and drug discovery, really needs a revolutionary application, or a whole new way of thinking about data, and we want to be a part of that.”
The Shape of Data
Ayasdi’s software is based on topological analysis, which visualizes relationships between data points as 2D shapes made up of interconnected nodes. The nodes represent the subjects under analysis: basketball players, say, or patients at a clinic. The most similar nodes will cluster together, with lines drawn between them, while highly dissimilar nodes will be split into separate shapes. The idea is that emergent patterns, like forks, loops, or separated clusters, will reveal meaningfully different subpopulations in the data.
To understand the program in action, consider a study that Lum has featured in detail on the Ayasdi Blog. Lum and her colleagues retrieved information on 870 tumors from The Cancer Genome Atlas (TCGA), an online database. The tumor samples came from cases of either breast or ovarian cancer, and included both genetic data and RNA expression levels. Lum’s team wanted to see if previous analyses of these tumors had missed any clinically relevant details about certain groups of cancer cases. They input each tumor into Ayasdi as a row, and each genetic characteristic as a column.
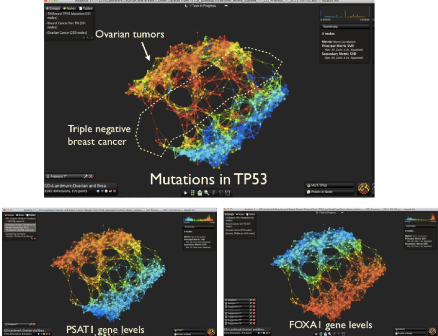
A topological network of 870 TCGA tumor samples generated in Ayasdi software. Three different views highlight mutations to TP53, and expression levels of two related genes. Image credit: Ayasdi
Ayasdi doesn’t use any preformed hypotheses in its analysis – Lum didn’t, for instance, tell the program in advance which tumors came from breast cancers, or ask it to arrange samples by the expression level of a particular gene. Instead, all the columns are sorted into “bins,” based on how closely they hang together. If the presence of a particular mutation to gene A correlates closely with expression levels of genes B and C, those three columns might be binned together. Bins can also overlap, so if the expression level of B also correlates with a mutation to gene D, then B’s expression may find itself in two bins.
The bins are a shortcut for independently associating every variable with every other variable, which makes the program very fast – even a very large dataset, like 870 tumors with thousands of genetic characteristics, can be resolved within seconds. “We can do up to a couple of million columns, sometimes in a couple of thousand patients,” Lum told Bio-IT World.
To create its visualizations, Ayasdi sorts the nodes into a 2D network based on their overall similarity, prioritizing the bins that best differentiate groups. At this point, with a basic topological map of the data, users can start asking more targeted questions.
“Once you have a network, you can ask what variables best explain a group of interest,” says Lum. “The software can run through the data automatically, and suggest features for you to select.” You can also layer metadata onto the network, coloring different nodes based on certain attributes. In the TCGA study, one important piece of metadata was which tumors came from breast cancer, and which from ovarian. Coloring those nodes revealed that Ayasdi had successfully separated the two types of cancer, using only the genetic data. Asking the program which features best distinguished between the breast and ovarian tumors also revealed a number of well-known oncogenes, giving important validation to the approach.
More uniquely, this particular study found a region of overlap between breast and ovarian cancers, which on closer inspection turned out to be triple-negative breast cancer cases, a clinical subgroup that has been especially difficult to treat. Probing the program on why it had placed these tumors in between the other breast cancers and the ovarian cancers, Lum’s team discovered that mutations to the tumor suppressor gene TP53 were very common in both ovarian and triple-negative breast cancer, although extremely rare in other breast cancers.
“People have suspected triple-negative may be a little different, but to show that TP53 mutations are pretty prevalent in triple-negative, which is very similar to the profile of ovarian cancer, is a very interesting finding,” says Lum. She also points out that the tumors in TCGA have already been studied exhaustively through traditional methods, without giving away this potentially actionable detail – future drug developers might now consider TP53 as a drug target for triple-negative breast cancer.
“Cancer is so heterogeneous that you might need to find more and more subpopulations to treat it more specifically,” Lum adds. “That’s why you need game-changing methods to get into it. If you use traditional methods that are more for looking at big signals, all the low-hanging fruits are basically gone.”
Built for Biology
Ayasdi has a number of clients in financial services, and is exploring partnerships in retail analysis. The company also has a large base of government contracts; much of the founders’ early research funding came from DARPA and the NSF, and big agencies like the FDA and CDC are now regular users. But the life sciences, and especially drug discovery, require a more specialized approach.
“There are many things specific to biology, like gene pathways, next-generation sequencing, compounds… which are very specific for people doing drug discovery, patient segmentation, and biomarker discovery,” says Lum. At the Bio-IT World Conference, Ayasdi unveiled both a new product and a major partnership to better tailor their platform to these use cases.
The product is Ayasdi Cure, which received a top prize at the conference’s Best of Show competition for new products. “Ayasdi Cure is an application on top of our core platform,” says Lum. “It is made for big pharma and life sciences, to tackle drug discovery from end to end.” The new application brings in tools that let users dig deeper into their raw data. A variant viewer lets users translate data on genetic variants from their topological maps to a Manhattan plot, quickly identifying the variants most closely associated with a trait of interest and showing where they occur in the genome. Ayasdi Cure also has built-in awareness of genetic pathways: rather than just telling users which individual genes are enriched or suppressed in a subpopulation, it can highlight an entire pathway whose expression levels are changed.
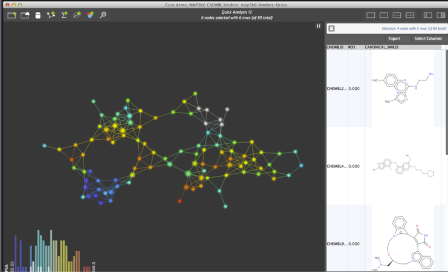
A network of compounds in Ayasdi Cure. On the right, the chemical structures of selected compounds are visualized onscreen. Image credit: Ayasdi
For drug developers, Ayasdi Cure also incorporates compound structures, derived from the public database ChEMBL and users’ proprietary datasets. “Any kind of compound databases will allow us to build a topological map of millions of compounds,” says Lum, “so you can quickly search for which compound violates the Lipinski rule, for example, or which compounds have great PK/PD.” Highlighting a region of the map will bring up the compounds’ structures, so chemists can inspect them manually.
“You can imagine this would be very useful all the way until you find your lead compound,” adds Lum. “And you still might want to go back and check how your lead compounds do against another database of compounds.” Pharmaceutical companies like Boehringer Ingelheim, which gave a collaborative presentation with Ayasdi at Bio-IT World, have already adopted Ayasdi Cure in their research pipelines.
Ayasdi’s second announcement at the conference was a partnership with CLC bio (now a part of QIAGEN), the world’s leading commercial provider of software for first-tier bioinformatics. CLC bio specializes in turning raw sequencing reads into useful data files, offering a software architecture that prepares samples for the more complex, functional type of analysis that takes place in Ayasdi Cure. “Because they have the structure already in place at CLC bio, and they also have partners that use them for mapping VCF files and getting sequences out, it’s a win-win solution for us,” says Lum. The two companies are now offering customers joint access to both CLC bio’s platforms and Ayasdi Cure.
“We add a lot of value and become an end-to-end solution,” Lum says.
Rethinking Common Diseases
While these new developments are targeted to basic research, some users are already moving Ayasdi’s platform into clinical settings. With unprecedented amounts of data on real patients from electronic medical records and next-generation sequencing, clinical programs are producing some of Ayasdi’s most exciting results and innovative projects.
At the Icahn School of Medicine at Mt. Sinai in New York City, Ayasdi has been deployed to rethink some of the most common chronic diseases in the U.S., asking if patients can be better sorted into subpopulations for more personalized treatment. Mt. Sinai has released the results of one study of Type 2 diabetes that answers this question with a resounding “yes!”
In a study of over 11,000 diabetic patients, Mt. Sinai’s Director of Biomedical Informatics Joel Dudley and his team input nearly 200,000 columns of both genetic markers and clinical data, such as blood levels and complicating symptoms, into the Ayasdi platform. Their topological analysis split patients into three major groups with little overlap: one with complications from kidney disease, one with cardiovascular and immune disorders, and one characterized by arterial disease and abnormal cholesterol levels. Representatives from both Mt. Sinai and Ayasdi have suggested that Type 2 diabetes may not be a meaningful clinical category at all; instead, patients might be better served by thinking about three entirely new types of diabetes.
“I think this is something that every researcher in diabetes has thought about, but it’s really hard to get to this answer,” says Lum. “I’m not surprised at all [by these results]. In fact, I think that if we increased the resolution*, we might even find more subpopulations. Diabetes is extremely heterogeneous. There are probably different co-morbidities that affect diabetes, and these patients probably respond to different kinds of medication, and maybe have different adverse events.”
This study won the Icahn School of Medicine the Editors’ Choice Award at the 2014 Bio-IT World Best Practices Awards, making Ayasdi the only company involved in top prizes in both the Best of Show and Best Practices competitions. Meanwhile, Joel Dudley’s team has been scaling up the Type 2 diabetes study to 30,000 patients, hoping to find underlying genetic variants or other biomarkers that could help rapidly sort patients into the appropriate categories, and possibly one day lead to new treatments. Mt. Sinai is also extending topological analysis to other chronic disease areas.
Ayasdi is hoping to pull off a similar trick with acute asthma. U-BIOPRED, an international consortium of academic partners, patient groups, and pharmaceutical companies, has been gathering very detailed data on a cohort of patients with asthma, including clinical histories and medications, but also gene expression in blood and lung cells and blood proteomics.
“Acute asthma is very much like diabetes,” says Lum, who has been recruited to the statistical advisory board of U-BIOPRED. “They know that there are probably many types of acute asthma, but the only way to [classify patients] in the clinic today is to blow into this apparatus, measure lung capacity, and say this person is mild, moderate, or severe. They know that’s not good enough.” Using Ayasdi’s platform, among other bioinformatics tools, U-BIOPRED wants to find subgroups that can be identified with blood markers and treated with unique medication regimens.
Pek Lum, Ayasdi's Vice President of Solutions and Chief Data Scientist. Image credit: Ayasdi
Ayasdi has also taken on some very novel sources of data. The Michael J. Fox Foundation recently partnered with Ayasdi to analyze data from a cell phone app distributed to people with Parkinson’s disease.
The Foundation was not aware of Ayasdi when it first undertook a trial study with the app, which collects audio data, gyroscopic information, and information from an accelerometer during the course of users’ everyday movements. In a limited pilot project, a small group of Parkinson’s patients and controls were given the app, and readings were recorded every second for a month. When Ayasdi heard about the project, they were excited to work with this little-explored longitudinal data, especially with such a high volume of data per subject.
“It’s a fairly new type of data, and it’s very, very noisy,” says Lum. “Imagine anyone carrying a cell phone – you could drop it, you could pick it up, you could go to bed or read a book. One of the challenges we talk about is how our platform approaches this kind of data.”
Ayasdi plotted each subject into a topological map based only on the data read out from the phones. “Not only did we manage to differentiate between controls and Parkinson’s, we think there are actually two subgroups,” says Lum. “Those could be mild and severe – we don’t know at this point, because we don’t know enough about the patients. But there are definitely two quite distinct subgroups of patients that have slightly different movements.” Ayasdi also discovered that their maps could be resolved using the accelerometer data alone, which, if the observation holds in larger studies, could make later research into patient movement easier. “You can imagine, in the future, if we do find that the accelerometer is the best, you can give them a very nice, small wearable device on their wrists or legs that only measures acceleration.”
Ayasdi is grateful for the experience working with longitudinal data, which could eventually become important in clinical practice. Mapping a patient’s movement over time might help to rapidly place them in an appropriate subgroup, by superimposing their motion data on a known topological map – and the same could hold true for other clinical measures. Lum imagines care centers collecting data from their patients in real time, constantly updating networks to find the best matches and treatments.
Lum and her colleagues think of Ayasdi’s analysis as fundamentally a new generation of bioinformatics. Traditional analysis, in her mind, has reached a sort of roadblock, where the limits are due less to the amount of available data, and more to an inability to find the right questions to ask. By visualizing data without hypotheses or suspected correlations, Ayasdi hopes to find the hidden routes through that barrier.
*One common technique for topological analysis with the Ayasdi software is to change the “resolution” of the networks. A lower resolution means that columns are sorted into larger bins, while at a higher resolution, the bins are made smaller. At the graphic level, this means that a higher resolution network will preserve only the strongest connections between nodes – groups that are more weakly associated tend to break apart as the bins are split. This can be used as quality control, to show only the strongest associations between variables at high resolutions, or only the most basic subpopulations at low resolutions.
In one creative project, Ayasdi has even used maps at different resolutions as a sort of data point in and of themselves. With the FDA, Ayasdi created a network of bacterial species based on their genetic data. Decrease the resolution, and you see more ancient taxonomic groups; gradually increase the resolution, and you start to see the evolutionary history of these species, as more and more closely related species break apart. In this way, Ayasdi built a phylogenetic tree of this set of bacteria.


