Illumina Opens BaseSpace to Proteomics Data
By Aaron Krol
October 8, 2014 | Another company with a 70% market share in its chosen field might have concluded by now that there are no more mountains to climb, but Illumina, the dominant player in genetic sequencing, is restless. The company is working furiously to consolidate an early lead in clinical sequencing with new FDA-cleared instruments and diagnostics. Meanwhile, the executive team has lately been stressing genetic data analysis as a key bottleneck to further expansion. BaseSpace, Illumina’s cloud-based data storage environment and informatics app store, is growing at a steady clip, as the company promotes the system as a place for innovators to put their analysis tools straight in the hands of researchers around the world.
Now, Illumina is making an early push into multi-omics, with the release of a series of apps for proteomic analysis in BaseSpace. Previously, BaseSpace has attracted apps for processing DNA and RNAseq data, but has not offered an infrastructure for related information like proteomic, metabolomic, or clinical and phenotypic data. The addition of mass spectrometry data to BaseSpace marks a first step toward supporting systems biology projects in Illumina’s own informatics environment.
The centerpiece of this initiative is a new partnership with AB SCIEX, whose lead line of mass spectrometers, the TripleTOF series, will now be able to deposit mass spec data into BaseSpace. As part of the collaboration, known as the OneOmics Project, AB SCIEX has also developed a set of four apps for turning mass spectra into protein profiles, and beginning to mine biological insights from the proteome.
“For longitudinal studies, BaseSpace’s ability to store both the raw data and the digitized proteome means you have all the data at your fingertips to share with collaborators,” Aaron Hudson, Senior Director of AB SCIEX’s Academic and Omics Business Unit, told Bio-IT World. “It takes all the headache out of the IT infrastructure.” Hudson adds that the cloud computing environment of BaseSpace will be a powerful enabler for users of AB SCIEX’s analytics software, saying that the resolution of mass spectra into a proteome, which would take three days on a desktop computer, can be done in BaseSpace in an hour.
Synergistic Technologies
The partnership between Illumina and AB SCIEX unites two companies who have spearheaded a drive toward higher throughputs and greater reproducibility in omics data collection. AB SCIEX’s analysis toolkit, says Jordan Stockton, Illumina’s Director of Marketing in Enterprise Informatics, “lends itself well to automation in the cloud, and has this aspect of being repeatable that other types of high-throughput molecular assays outside of genomics just don’t have.”
AB SCIEX is the creator of the SWATH technique for resolving mass spectra into their constituent proteins. SWATH, released in 2012, was one of the earliest and most comprehensive commercial examples of data-independent acquisition (DIA) mass spec. Traditional, data-dependent mass spec makes two scans of each sample, using a preliminary scan as a filter to decide which data to include and exclude. By contrast, DIA methods take advantage of faster mass spec instruments that can capture the full range of protein signatures without discarding data. SWATH offers reproducible data collection and analysis, appropriate for the kinds of wide-ranging projects Illumina hopes to foster in BaseSpace.
The four apps AB SCIEX has developed for the OneOmics Project are unique to BaseSpace, and have a biological approach in mind. Protein Expression Extractor translates mass spec data into a proteome; Protein Expression Assembler is used to quantify changes in protein expression; and Protein Expression Analytics gives bioinformaticians a window into the SWATH analytics parameters for quality control.
The fourth app, Protein Expression Browser, will be especially valuable for uniting proteomic data with other biological information. The app gives users a project-wide view of protein expression in different samples, and allows them to drill into specific proteins for knowledge about their cellular pathways, genetic precursors, and the extent of their up- or down-regulation.
“I don’t think proteomics has seen anything like it just yet,” says Hudson. “It’s a tool that really gives a great first analysis of what’s happening with the proteins in the cell.”
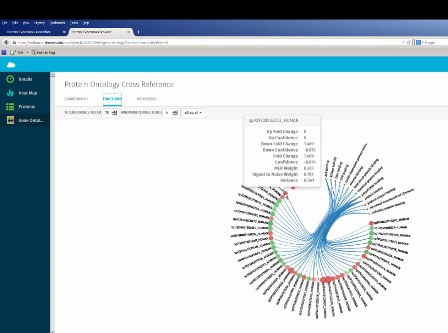
A screenshot from Protein Expression Browser, showing the gene ontology view with protein expression cross-referenced against related molecular functions. Image credit: AB SCIEX
Both Illumina and AB SCIEX expect users to combine the SWATH tools in BaseSpace with RNAseq analysis apps, to discover how the transcriptome, the set of RNA molecules transcribed from DNA in a given cell type, is translated into proteins.
“There’s been a long-held belief that only a small fraction of the mature transcripts are translated, and now we have the ability to see which transcripts are expressed on the protein level,” says Stockton. “This is a tool for uncovering mechanistic insights about pathways we think we already understand. It may uncover new isoforms of proteins that interact in ways we don’t know.” On the medical side, the OneOmics project will also offer a useful toolkit for discovering new biomarkers that could be used to diagnose or subtype disease.
The BaseSpace Connection
While AB SCIEX is the major partner in the OneOmics Project, other organizations are also creating new proteomics apps for BaseSpace, designed to work with TripleTOF and SWATH data. An app from Advaita Bio combines both proteome and transcriptome data to visually represent the up- and down-regulation of cellular pathways, as a natural extension of the Protein Expression Browser. The Institute for Systems Biology in Seattle has also contributed an app, providing spectral libraries to help SWATH identify proteins in samples from different species. The initial release includes libraries for human, the tuberculosis bacterium, and Saccharomyces cerevisiae, a species of yeast commonly used as a model organism.
Yale University is working on another app that will convert experimental RNAseq data into predictive proteomics data, allowing users to move straight from genetic sequencing to proteome analysis. The OneOmics partners expect more app developers to enter the BaseSpace proteomics ecosystem as this first set of tools is adopted by users. “I think what you’ll see is people start to innovate on this in ways we never thought possible,” says Hudson.
The
AB SCIEX apps are currently in beta release, and are free to use, although the
company does eventually plan to charge on either a per-use or subscription
basis. Illumina is also offering free storage of mass spec data for the time
being, as the company mulls over its eventual cloud storage model.


