Nanopore Sequencing Is Here to Stay
By Aaron Krol
December 22, 2014 | The black boxes from Oxford Nanopore Technologies arrive in the mail in FedEx envelopes, looking a bit like individually packaged bars of soap. Oxford Nanopore started shipping its first products this April, and the company has gone to some trouble to make the packaging feel like any other consumer electronic. An early access customer opening one of the boxes will find a molded plastic inlay, a carefully folded USB cable, and a device called the MinION, a three-inch-long gadget with flashing LED lights in three colors.
The lights might strike some users as a little hokey, giving the instrument the vibe of a prop from a ‘60s sci fi movie. Then again, that’s a pretty fair description of the MinION, a DNA sequencer so small it could easily be confused with a thumb drive or an off-brand mp3 player.
To Nick Loman, a microbial geneticist at the University of Birmingham, picking up his first MinION was a slightly surreal experience. Loman’s lab started doing its own gene sequencing in 2009, with an instrument called the Genome Sequencer FLX. It looked like a waist-high file cabinet with a large office printer sitting on top, and just five years ago, it was the bleeding edge of genomics. And while most sequencers these days dispense with the file cabinet (an onboard computer), they’re still bulky enough to make their presence known: things you’d probably call instruments, and almost certainly not gizmos.
“The benchtop sequencers opened up the market to a certain degree,” says Loman. “You started seeing [them] in intensive research groups, and in the clinic. But what if anyone could have this hanging off their key ring, and go do sequencing? That’s an insane idea, and we don’t really know what it’s going to mean in terms of the potential applications. We’re very much at the start of thinking about what we might be able to do, if anyone can just sequence anything, anywhere they are.”
The MinION is a radical step in the evolution of reading DNA, a smartphone in a world of supercomputers. Its boosters have started talking about the whole science of genetics as if it could be more like a hobby. They want to take MinIONs to remote clinics and outbreak zones, into sewers to track bacteria and viruses, out to nature reserves to study ecological changes or hunt for unknown species — even to ports and border stations as an enforcement device for catching poachers.
Until recently, these ideas would have seemed pretty far-fetched. It’s not just that DNA sequencers are big machines, or that running them is an elaborate (and expensive!) process. It’s not just that dealing with the data they produce is a job for dedicated experts, who use gigabytes of memory and days of compute time to wring useful information out of their raw DNA sequence.
It’s that Oxford Nanopore specifically looked a little dubious. After strongly hinting in early 2012 that a product release was just around the corner, they all but dropped off the map — except to announce occasional rounds of funding, eventually totaling over $280 million. They rarely spoke to the press (and declined to comment for this story). Clive Brown, the CTO and often the public face of the company, would pop up to show impressive results from mysterious prototypes, which no one could verify. Meanwhile, a number of biologists argued that the technology — utterly different than any other sequencer on the market — could never be made to work as promised.
Even after the first instruments started shipping this year, for a limited test release called the MinION Access Program (MAP), lingering doubts have hovered over the project. MAP participants have to test their devices on a pre-specified sample, and can’t share any data with the public until they certify to Oxford Nanopore that the instrument is “acceptable.” No one has matched the data quality the company claims to see internally. Yet, eight months after those first boxes were torn open, hundreds of MinIONs have now been shipped to roughly 40 countries, and users are starting to speak up about what the device can do. There’s not much data out there, not yet, but it’s enough to say three things.
Oxford Nanopore has made a real sequencer. It can do real science. And that is a really big deal.
The DNA Thread
The idea of a nanopore sequencer has been floating around since 1989, the brainchild of a biophysicist from UC-Santa Cruz named David Deamer. Deamer’s insight was to take the chemistry problem of creating molecules that can manipulate DNA dissolved in liquid, and turn it into an engineering problem of catching the DNA in a single, predictable place. That place is a “nanopore,” a hole so small that only a single molecule of DNA can fit through.
By suspending lots of nanopores in a liquid charged with an electrical current, Deamer believed it should be possible to read DNA as it was threaded through the pores. Each DNA base, the “letters” of the DNA code, would disrupt the current in a signature way when it passed through, and that signal could be read by a computer to decode the whole sequence.
 Of course, there were a few kinks to work out. Like, for
instance, the fact that no one had ever made a nanopore small enough for the
design. Or the problem of actually getting the DNA where it needed to go: it’s
all very well to say a DNA molecule is threaded through a nanopore, but when
you consider that the hole, at one nanometer wide, is a million times narrower
than the head of a pin, the sewing metaphor pretty much falls apart.
Of course, there were a few kinks to work out. Like, for
instance, the fact that no one had ever made a nanopore small enough for the
design. Or the problem of actually getting the DNA where it needed to go: it’s
all very well to say a DNA molecule is threaded through a nanopore, but when
you consider that the hole, at one nanometer wide, is a million times narrower
than the head of a pin, the sewing metaphor pretty much falls apart.
“There have been skeptics all along who said that nanopore sequencing couldn’t possibly work,” says Mark Akeson, a colleague of Deamer’s at UCSC who has been wrestling with these issues for more than 15 years. But he adds that no one could quite say that a nanopore sequencer was physically impossible to build. “Instead, the response tended to be, ‘well, it's just too hard!’”
Bit by bit, Deamer and other believers started to cobble together prototypes in their academic labs. In an important boost, the National Institutes of Health invested heavily in nanopore sequencing through the Advanced DNA Sequencing Technologies grants, a program that has had a hand in almost all the biggest sequencing breakthroughs of the past ten years. Akeson received millions in funding through that program, and made a crucial advance in 2010 with his colleague Kate Lieberman, showing that a particular DNA polymerase, attached to a nanopore made of protein, could attract DNA molecules to the pore and guide them through at a controlled speed.
Still, the systems built for these experiments were jury-rigged, imprecise, and needed the careful attention of their creators to work at all. Oxford Nanopore promised more. Hoping that Oxford’s deep pockets and large scientific staff could make nanopore sequencing a reality, centers like UCSC licensed key patents to the company. (In addition to being the inventor of several patents used by Oxford Nanopore, Akeson is also a shareholder.)
Today, using his MinIONs in his Santa Cruz lab, Akeson is convinced that faith has paid off. Academics proved a nanopore sequencer could be built in principle, he says, but “everything from that point on was implemented by ONT.”
The MinION largely builds on the experimental work at UCSC: it uses similar protein nanopores, and a similar polymerase. But years of painstaking refinements have allowed the instrument to perform a delicate chemical dance, in which double-stranded DNA is unzipped, tied together at one end, and guided gently through the nanopore as a single string. This dance occurs hundreds of times in concert, at such a careful pace that tiny changes to the voltage inside the MinION can be picked up and fed to a computer in real time.
“Think about it,” says Akeson. “They have engineered a 100-gram device, composed of 500 pores each with its own dedicated amplifier, that reads bases at Angstrom precision tens of thousands of times in a row — and it is shipped by FedEx to anywhere on the globe!”
Hunting Bugs
“It’s a minor miracle that it worked out of the box,” says Loman.
When his team at Birmingham got its first MinION this April,
they scrambled to try it out on whatever sample happened to be in the fridge. The
lucky bacterium was Pseudomonas
aeruginosa, which the lab was studying in connection with hospital
outbreaks. “The very first few experiments we did, we just wanted to see
whether [the MinION] generated any sequence that was recognizably Pseudomonas,” says Loman. They put some
DNA and necessary reagents into the sequencer, plugged it into a laptop, and booted
up the software provided by Oxford Nanopore, a cloud-based platform called MinKNOW.
When MinKNOW gave back sequence, they ran a second program called BLAST, which can
quickly compare a DNA read with known genomes and hunt for matches.
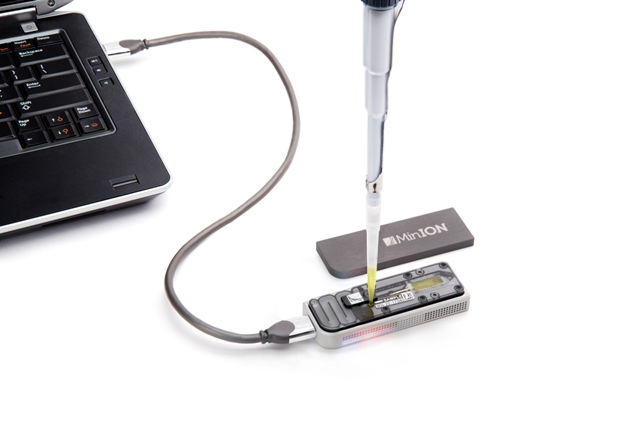

The MinION, close up. Image credit: Oxford Nanopore Technologies
The first few hundred reads didn’t look like anything in particular — the reads were short and the sequence questionable, as if the MinION were still getting its bearings. “And then we found one that was about eight kilobases long, which is extremely long for any sequencer,” says Loman. “[BLAST mapped it] across the entire length of a part of the Pseudomonas genome that tells you what its serotype is, which is an important bit of information from a clinical microbiology point of view.”
Loman posted this read on Figshare, a website where scientists publish small pieces of data. It was the first MinION read made public by anyone other than Oxford Nanopore, and before long, it had become one of the 25 most-viewed publications on the site. Loman’s P. aeruginosa read showed that the sequencer was not only running, but able to pick up valuable information. If reads like this one could be achieved in a hospital, the staff could use MinIONs to learn the species and serotypes of bacteria causing undiagnosed infections.
Every MAP member I spoke with brought up the subject of taking MinIONs to the clinic. Care centers without sophisticated labs could one day use the device as a portable, nearly all-purpose diagnostic for infectious disease — and its DNA data could turn up crucial details that other tests miss, like which bacteria have mutations that make them resistant to antibiotics. “There’s very little in the way of setup required,” says Loman. “You can imagine throwing one in a suitcase and taking it down to Sierra Leone to look at the Ebola outbreak.”
This is more than just speculation. Loman and his colleagues have already used the MinION during a local hospital outbreak of Salmonella, to test whether the device could tell the difference between the hospital’s bacteria and other samples from an unrelated strain. “Very rapidly, within an hour or two, we can get a firm assignation to a strain either being part of the outbreak or not,” says Loman, a result he confirmed by comparison with another sequencer, the MiSeq sold by market leader Illumina.
“It’s down to us, the researchers, to figure out the cool stuff we can do with it,” he adds. “I think it’s quite clear that sequencing in three years’ time is going to look quite different from how it does today.”
Do-It-Yourself Sequencing
To some MAP members, using the MinION for clinical sequencing seems almost old hat. After all, high-end medical centers are slowly embracing the genomics revolution, and it’s probably only a matter of time before sequencing takes off in disease surveillance. The really interesting question is whether Oxford Nanopore can do for genetics what Kodak did for photography, creating an “amateur” movement that changes the very nature of the art.
“We’re looking at a complete transformation of how people use sequence data, and under what conditions, and who will be using it,” says Brook Milligan, a MAP participant at New Mexico State University. “People who up to now have thought that genetics is not relevant, or beyond their reach, will be empowered to use it, and that creates all kinds of opportunities.”
Milligan is director of the Conservation Genomics Lab (CGL), an institution that has already benefited from seismic shifts in genetics. Ten years ago, “conservation genomics” would have been almost unimaginable: the field depends on sequencing DNA quickly enough, and in such quantities, that it can form an almost real-time picture of a wild population’s health, size, and distribution.
With much cheaper, faster sequencing machines, it’s become possible to apply genetics to a host of unexpected fields, and today, members of the CGL and its partners can be found collecting samples out in the wild to monitor populations of checkerspot butterflies, California poppies, and other at-risk species. “[We’re] answering questions like, how many individuals are there in a population you can’t necessarily see?” says Milligan. “What habitats are they using? What are their movement patterns?”
Tackling these questions relies on collecting lots of reference material, the “background noise” of a population’s genetic variety, and that involves a lot of time in the field. If Milligan’s team could bring a few MinIONs on their expeditions, specimens wouldn’t have to be shipped back to the lab for processing.

The MinION isn’t quite point-and-click. Like any sequencer, it needs to be loaded with isolated DNA molecules, which are not exactly lying around waiting to be scooped up. You need at least two big pieces of lab equipment to prepare samples for the MinION: a centrifuge to isolate your DNA, and cold storage for the enzymes Oxford Nanopore provides to guide DNA through the pores. (For long-term storage the reagents should be frozen, but they will survive for a while at 4° C.)
Still, compared to most benchtop sequencers, this isn’t a huge chore. Most importantly, users don’t have to make dozens of copies of their DNA to get a signal, a time-consuming process called polymerase chain reaction. Without PCR, nearly all the lab work can be done at room temperature, and some MAP members have reported getting their samples ready in an hour or less. Future versions of the MinION could be even simpler to use: Oxford Nanopore is working on a one-step process using a specially-engineered transposase, an enzyme that cuts up and transports DNA.
Milligan believes the prep work could easily be done in a makeshift lab in any indoor space with electricity, or even out on a boat. He brings up the DIY biology community, a small but growing movement of hobbyists who have, among other things, designed 3D-printed centrifuge rotors that attach to ordinary power tools. Add a mini fridge and a laptop, and you have a primitive nanopore sequencing station.
“It would be hard to do it literally in a tent,” Milligan says, before pausing to think it over. “[But] at least under certain circumstances, we probably could.”
Milligan’s pet passion is the trade in illegal biologics: poaching, logging in protected forests, fishing of endangered species. “Genetic information could be used to track that — the flow, and where it’s coming from, and what populations are being targeted,” he says. Right now this is highly impractical, because samples can’t be sequenced where they’re found. But Milligan believes that, with MinIONs deployed at ports, or border stations, or on board Coast Guard ships, any suspect biologics could be tested on the spot, and seized if necessary. With good enough reference material, it should be possible to trace not just the species of a fish or timber sample, but exactly where it came from.
With a lot of the burden of lab work removed, Milligan thinks the fundamental skills needed to work with genetics are poised to change. If, someday soon, anyone could be equipped to sequence DNA, the real challenge will be figuring out what to do with that data once we have it.
Signal and Noise
Using a MinION is startlingly like using any other USB-connected gadget. Plug it into your laptop, run MinKNOW, and wait. You’re even treated to a live view of the “squiggle plot,” a graph of the electrical signal from the pores, with the voltage moving up and down as DNA passes through.
Unlike when you connect a digital camera to your computer, however, the backend work being performed is massive. The MinION’s sensors take tens of thousands of reads a second, so vast amounts of data are being processed in those squiggle plots. That gets compressed into a much simpler graph in real time, and inside the Oxford Nanopore cloud, changes in the electrical current are translated into DNA bases, in a common data file format called HDF5.
All the really interesting information comes from mining those HDF5 files. It’s no small task. A whole professional class of bioinformaticians exists to build programs that turn bland strings of A’s, T’s, C’s and G’s into real-world knowledge — and because the MinION is profoundly different from other sequencers, many of their favorite tools work poorly or not at all with its data.
Some of the MinION’s quirks are very much to its advantage. For example, nearly every other sequencer splits DNA into tiny fragments, less than 500 bases long. The MinION will read essentially any size strand of DNA you give it; Oxford Nanopore recommends 8,000 bases as a good median length, but in private, some MAP members have claimed to see validated reads as long as 100,000 bases.
Long reads are a major asset, because the process of
stitching short reads together tends to lose information. In highly repetitive regions
of the genome, it can be impossible to figure out the correct order of short
reads — a limitation that leaves gaps in even the best-studied genomes. One of
Mark Akeson’s hopes for nanopore sequencing is to fill in parts of the human
genome that have never been sequenced.
Other features of the MinION’s reads are more problematic. Above all, high error rates have haunted Oxford Nanopore since the company first started releasing data. In many ways, the entire MAP has been a relentless race to push the MinION’s error rate down to reasonable levels, with multiple replacements of the hardware, software, and chemistry all devoted to controlling reads that still call the wrong DNA bases as much as 30% of the time. Filtering out the worst reads can help, and the latest version of MinKNOW actually throws away more reads than it keeps — either because only one strand of the double-stranded DNA made it through the pore, or because the DNA passed through too quickly to give a clear signal. Nonetheless, few if any MAP members have achieved the roughly 95% accuracy Oxford Nanopore claims to see internally.
Even worse, the MinION’s errors are “biased,” consistently misinterpreting the same changes to the electrical signal. That means running the instrument multiple times on a sample will tend to produce the same mistakes again, instead of correcting errors made the first time. Its errors also seem to cluster, sometimes producing long strings of gibberish where one bad base call created a domino effect down the line.
A few existing tools built for Pacific Biosciences, another company with long reads and high error rates, do seem to play fairly well with the MinION — but for the most part, early users are either rewriting their workflows to deal with nanopore data, or building new ones from scratch. Nick Loman is co-author of one of the most popular of these releases, a toolkit called Poretools.
“[Oxford Nanopore] are trying to encourage a diverse user community, and that includes people without a huge amount of bioinformatics experience,” says Loman. “So there was quite a lot of demand for some easy toolkits.” And certainly, recreating the robust set of tools available for, say, Illumina’s sequencers is a top priority.
But there’s also a chance to reconsider how bioinformatics is done. Up to now, a complex tangle of homebrewed tools and popular open source pipelines has worked well enough for genetics, if only because getting the raw DNA data to analyze was fairly complicated in its own right. If more casual users start working with sequencers, however, they’re going to need a large body of purpose-built tools that can be run with a single click.
DNA Coders
More than with any other sequencer, the idea of “apps” feels relevant to the MinION. Users might easily be amateurs with little or no experience with DNA. Supporting them will be a legion of programmers whose job is to find the fastest way to turn raw signal into something a hospital worker, government agent, or even hobbyist would want to know about a biological sample.
To pave the way for fieldwork with the MinION, Brook Milligan has been working on species identification, testing bacterial samples to see how well he can match them to known genomes. Despite the MinION’s high error rates, he says, his pipeline can reliably tell apart strains of bacteria that are 99.998% genetically identical — one difference for every 50,000 bases — within 30 minutes of running the sequencer.
At The Genome Analysis Centre in Norwich, another MAP participant, Matt Clark, has been trying to achieve the sequencing equivalent of a speed run. Clark’s program, a nanopore-specific software tool called Kontaminant 2, pulls 21-letter stretches of DNA (or “21-mers”) out of the MinION’s much longer reads. Using these short snippets, Clark can swiftly make species matches without going to the trouble of aligning entire reads to reference genomes. “The advantage of doing it via k-mers is it’s incredibly fast,” says Clark. “It’s the kind of thing you could do on a phone. If you run it on a laptop, it takes seconds.”
Tools like Kontaminant 2 prove that the MinION’s error rate isn’t a fatal flaw for the device. “It has quite long stretches of perfect sequence,” says Clark, even if it sometimes goes off the rails. That means shortcuts like k-mer extraction can get very high quality sequence out of the MinION, good enough to use the sequencer on its own in the field. Many of the most exciting applications, after all, are at heart no more than matching puzzles, drawing lines between a DNA sample and a strain of virus, antibiotic resistance gene, or a genetic marker that proves a shipment of bluefin tuna came from protected waters.
In the future, the algorithms that do this sophisticated cross-stitching could be placed in the cloud alongside MinKNOW, giving almost unlimited compute power to field teams working off a single laptop. Alternatively, Oxford Nanopore could release its base calling software for download, letting scientists run all the computations on their own computers. “A lot of it is about speed,” says Clark. “If you have to send things up to the cloud,” then you have to wait for a response from MinKNOW to run your own tools. A version of MinKNOW on your own hard drive, on the other hand, could start toying with data even before the MinION finishes reading the molecule. Since the raw signal from the MinION is already stored locally — so users aren’t completely dependent on their Internet connections — it would be a small step to read the squiggle plots locally, too.
Whether nanopore programs end up in the cloud or on laptops, shifting the action to the IT side lets a much larger developer community get involved in DNA analysis. “We have millions of people around the world working on open source software,” says Milligan. “We don’t have millions of people working on molecular genetics… The laboratory tasks are going to be trivial, and the analysis is going to be much more complicated and exciting.”
The Future of Genetics
Oxford Nanopore is still going through some teething troubles. The company has been making rapid changes throughout the MAP, mailing out new instruments and chemistry kits, and quality control has lagged a bit behind. The MinION flow cells, the cartridges where samples are inserted, are of notoriously uneven quality: while each cell contains 512 nanopores, even in the best case fewer than half of those provide a good signal. In the worst case, users just have to wait for a new shipment to work with the instrument at all. One MAP member described it as playing “flow cell roulette.”
There’s also a price to pay for miniaturization. The MinION generates small amounts of data compared with other sequencers — enough to work with bacteria or yeasts, but not the human genome, at least not without some clever lab work to narrow down your DNA samples to one or two regions of interest. “We’d be interested in doing more ambitious things, and I think the yield isn’t really there,” says Clark. And the error rate will continue to be a drag on many experiments: it would be irresponsible, for example, to use the MinION to sequence brand new genomes, unless its reads were combined with a second instrument to check the accuracy. On the other hand, Akeson, whose work includes using machine learning to teach computers to read nanopore data, is optimistic that the errors are basically a software problem, which could be fixed just by reinterpreting the squiggle plots.
 The MinION may not be an engine for new discoveries, but for
the first time in the history of genetics, the frontiers of the science don’t
appear to be cracking the next species, or even sequencing human genomes on the
cheap. The future of genetics may well be about access, and if that’s the case,
a lot of the commercial and regulatory systems around the science have some
catching up to do.
The MinION may not be an engine for new discoveries, but for
the first time in the history of genetics, the frontiers of the science don’t
appear to be cracking the next species, or even sequencing human genomes on the
cheap. The future of genetics may well be about access, and if that’s the case,
a lot of the commercial and regulatory systems around the science have some
catching up to do.
Insiders have speculated for years about a world where everyone could be a consumer of genetic information — where we could be sequenced at birth, when we get sick, when we decide to have children. Few have wondered what happens when we can all be producers of DNA data, because the idea seemed so far out of reach. While sequencers exist today that can read a human genome for a few thousand dollars, the machines themselves cost six figures or more, making them the almost exclusive province of a few major sequencing centers.
MAP members, however, are buying their MinIONs for $1,000 each, and Oxford Nanopore has teased that the final price could be even less for certain customers. Many labs, even smaller ones, are scooping up several of the devices, which is unprecedented for a sequencer still in active development. (Loman, a self-described fanboy, has “five or six.”) When the MinION is finally released for general sale, it will be priced less like an MRI machine than a nice telescope — and sequencing will be a little less like a medical procedure, a little more like stargazing.
That doesn’t mean we’re heading for the world of GATTACA, where you have to watch every hair and skin cell you leave behind. But it does erode the idea that DNA is some foreign substance observable only in a sterile lab, like biology’s own Higgs boson. DNA has always been all around us, and in a profound way, it’s closer now than ever before. With a pocketsize sequencer, it’s time to imagine a world where genetic data could be as near to hand as your cell phone, as ready-to-go as a ride from Lyft.
That’s an insane idea, and we don’t really know what it’s going to mean.
Illustrations: Aaron Krol
Works Referenced and Further Reading
Akeson and Lieberman's paper demonstrating a key step toward nanopore sequencing: "Processive Replication of Single DNA Molecules in a Nanopore Catalyzed by phi29 DNA Polymerase." Kate Lieberman et al. Journal of the American Chemical Society, 2010, 132(50).
Nick Loman's Poretools: "Poretools: a toolkit for analyzing nanopore sequence data." Nicholas Loman and Aaron Quinlan. Bioinformatics, Oxford University Press, 2014.
A whole bacterial genome sequenced using the MinION: "A reference bacterial genome dataset generated on the MinION portable single-molecule nanopore sequencer." Joshua Quick, Aaron Quinlan and Nicholas Loman. Gigascience, 2014, 3:22.
An antibiotic resistance gene placed in context using the MinION: "MinION nanopore sequencing identifies the position and structure of a bacterial antibiotic resistance island." Philip Ashton et al. Nature Biotechnology, 2014.
Bio-IT World's two previous features on Oxford Nanopore, in 2009 and 2012: "Brown & Oxford Nanopore," "Oxford Strikes First in DNA Sequencing Nanopore Wars."


