The Tsetse Fly and the Test for Genomics in Africa
By Aaron Krol
August 6, 2015 | At the Biotechnology Research Institute in Muguga, Kenya, about 15 miles west of Nairobi, Paul Mireji is trying to figure out how tsetse flies find their prey. The half-inch flies are hyperactive bloodsuckers, who will take a blood meal once a day if they can get it — and that makes them ideal vectors for any parasites who want to hitch a ride. Their passengers include a protozoan called Trypanosoma brucei, the microscopic agent of Africa’s most unique public health threat.
T. brucei has the rare ability to pass from the bloodstream into the brain, where it causes a profound neurological disease called African trypanosomiasis, better known as sleeping sickness after the extreme cycles of sleeping and wakefulness that are common symptoms. Spread easily by its voracious insect hosts, T. brucei infects and kills thousands of people every year ― and that’s a small fraction of its toll during a major epidemic, like the one that roiled across more than 30 countries for almost three decades beginning in 1970. There is no vaccine, and the few drugs are almost as bad as the disease itself: one, reserved for the latest stages of trypanosomiasis, is a compound of arsenic that poisons the patient and parasite alike.
“Suppression of tsetse flies is the most promising route to trypanosomiasis control,” Mireji explains, speaking through a webcam from his lab at the BRI. “We have been able to develop some very good tools for control of the animal form of trypanosomiasis, but we still don’t have tools for better control of the species of tsetse fly that spread the human form.”
Muguga is not far from the tsetse fly’s natural habitat, a two-thousand-mile-wide belt of woodlands and open brush stretching across the center of the continent from the Atlantic coast to the Indian Ocean. The BRI is home to a colony of flies collected from the nearby Kenyan countryside, and part of Mireji’s experiments involves putting flies through a wind tunnel to see how they respond to different scent cues.
But like many modern biologists, Mireji does much of his work at the computer. There, he crawls through the genetic code of the fly, looking for genes whose role in behavior can be guessed at by comparing them to genes in related species like fruit flies. Mireji was one of the first scientists to identify genes involved in the tsetse fly’s senses of smell and taste, discovering that it likely responds to a much narrower range of molecules than its relatives. The next step is creating strains of the fly where certain of these genes are shut off, to learn how those changes will affect behavior.
“We still don’t understand the olfactory system of these tsetse flies that they use to locate their hosts,” Mireji says. “We are trying to see whether, using our knowledge of tsetse fly genomics, we can find molecules that repel the fly and molecules that attract the fly.” That information could lead to more effective traps and repellants, stopping the flies before they can unload their microscopic hitchhikers.
Mireji’s work at the BRI is just one local offshoot of a global project involving scientists from 18 countries across Africa and more in the United States, Japan, and Europe, who worked together for more than a decade to produce the first whole genome sequence of a tsetse fly species, Glossina morsitans. Though it was slow going by the standards of genomics today, the International Glossina Genome Initiative (IGGI), which announced its success in April 2014, put African minds to work on a distinctly African problem, allowing scientists to reach a more intimate understanding of a species that has caused deadly epidemics across an entire continent. Along the way, investments in local research centers, and large training courses in the computational techniques needed to investigate the genome, ensured that the African members of the IGGI will be able to continue their work with or without the help of their big international funders.
“Ultimately, it helped in building infrastructure that we can use,” says Mireji. “We can take advantage of being among people who are directly affected by the disease. We have the real tsetse flies, real parasites, and we want to see how we can translate bioinformatics into the field.”
A Bizarre Foe
The tsetse fly has traced a winding path from its house-fly-like ancestors.
“There were lots of interesting reasons to sequence this genome besides that it’s a vector,” says Serap Aksoy, one of the organizers of the IGGI. Aksoy has worked on the genetics of trypanosomes since the 1980s, and her lab at Yale School of Public Health maintains a colony that is surely one of the largest concentrations of tsetse flies outsides of Africa. “Biologically,” she says of the fly she has studied for 30 years, “it has many unique characteristics.”
Tsetse flies are extreme specialists, getting all their energy from blood ― unlike their distant relatives mosquitoes, which are nectar eaters first and bloodsuckers second. That dependence on blood has shaped the fly’s entire lifecycle and anatomy. To survive on this single resource, tsetse flies have evolved to give special care to their young, who are entirely reliant on the mother to convert her blood meals into the full range of nutrients they will need to make it to adulthood.
The result is an insect with a life uncannily like a mammal’s. A pregnant tsetse fly does not lay a batch of eggs like a house fly, but nurses a single larva at a time in her uterus. The developing larva drinks a nutrient-rich “milk” given out by a unique gland inside the uterine wall. After a weeklong gestation, the mother gives live birth, and her now independent larva prepares for its metamorphosis into an adult, needing no more meals until it’s ready to take blood on its own.

This photo of a pregnant tsetse fly shows the swollen abdomen where the fly is producing its milk and nursing a single larva. Image credit: Geoffrey Attardo, Yale School of Public Health
Exploring tsetse genetics has deepened our knowledge of every phase of this lifecycle. For instance, it was known that tsetse flies have a unique population of gut bacteria that help them process blood into a lifelong food source; now we realize that these bacteria have profoundly stunted genomes, and have probably co-evolved with their hosts to the point that they are completely dependent on the tsetse gut environment. We’ve also learned just how much of a female tsetse fly’s energy goes to producing her milk. RNA studies reveal that as much as half of the RNA translating a pregnant tsetse fly’s genes into proteins is devoted to this single task.
Any of these insights might be turned into weapons against the spread of trypanosomiasis. “The genome identified all of these novel genes that produce proteins unique to the milk,” says Aksoy. “This is a perfect opportunity to interfere.” Other promising targets are the chemosensitivity genes that tsetse flies use to navigate, and the immune genes that affect their relationships with trypanosomes.
The current bounty of leads in the battle with sleeping sickness, however, masks a long period of neglect. Although there was a catastrophic epidemic in the 1970s, ‘80s, and ‘90s, the main drug regimens for treating trypanosomiasis have not been updated since 1949. The drugs that do exist, some of them almost a century old, come with devastating side effects ― and the parasites are starting to develop resistance to them. Funding for new therapies, or to create a vaccine, mostly dried up with colonialism, despite efforts by a few dedicated organizations like the Swiss Tropical Institute to renew focus on the disease.
By the turn of the century, the scientists still confronting trypanosomiasis were ready to start afresh. “Around that time, we had the emergence of new technologies in genomics that were revolutionizing our understanding in many systems,” Aksoy remembers. “If we were going to do something novel in this field, expanding our genetic knowledge of tsetse and tsetse trypanosome biology would be able to take us to that next level.” Aksoy and other scientists, many of whom met through related work at the World Health Organization (WHO), proposed a project to sequence the genome of a tsetse fly.
In brainstorming sessions at the WHO, this nascent team concluded that, although reading DNA could happen anywhere, the effort to understand the tsetse genome would have to take place in Africa to have the best chance of translating genetic information into actual tools for controlling trypanosomiasis. Fortunately, the South African government was then investing heavily in computational biology through the South African National Bioinformatics Institute (SANBI) in Cape Town, providing a natural home base for genomics operations in Africa. In 2004, the WHO pulled together scientists from SANBI, Yale, and other international centers for a meeting in Geneva, the first convention of the International Glossina Genome Initiative.
Grant by Grant
At the time meetings of IGGI members began, sequencing the genome of a complex organism like a tsetse fly was an elaborate process. Only a few research institutes in the world had the equipment to embark on a whole genome sequence ― and to even begin, they would need the DNA of the fly to be broken in small units and cloned into bacteria.
With only a small grant from the WHO, a task that large was beyond the IGGI’s reach. Still, having a reliable, ten-year source of funding was a crucial springboard. “The funds we received were surely not enough to do much in terms of the genomics, but they allowed us to bring a group together annually,” says Aksoy. “It allowed us to work as a community to look for money for the big project, and in the meantime it allowed us to do small trainings and workshops to get our feet wet. It kept the community alive.”
From the outset, the IGGI acted as though it would eventually have the resources to do an entire genome, believing that donors would be attracted to a big goal. Members discussed which species of tsetse fly to go after: G. palpalis, which is more active in West Africa and causes the large majority of trypanosomiasis cases, or G. morsitans, most active in East Africa where a rarer but more virulent form of the disease is found. After settling on morsitans, the IGGI had to cultivate a colony of flies to provide the DNA.
“It’s a difficult fly to work with,” says Aksoy. The tsetse’s slow reproductive cycle is a problem when you’re trying to collect a lot of DNA from closely related flies — and the intensive but highly accurate “Sanger sequencing” method used for the project required a lot of DNA. “We had to make DNA from one mom and her offspring,” Aksoy says, “so that we reduced the heterogeneity in the final product. It was an experiment to see if we could maintain this single female line.”
Meanwhile, with help from the WHO and the Fogarty International Center, the IGGI invested in workshops and training sessions for scientists at centers like Gulu University in Uganda and the BRI in Kenya (at that time called the Kenya Agricultural Research Institute). The trainings, some held at SANBI and some through overseas exchanges, entrenched the project as a local effort and made sure scientists in participating countries would have the skills and knowledge to scale up with new funding. Aksoy recalls that a number of participants in those first workshops went on to be co-authors of the paper presenting the G. morsitans genome almost ten years later. Some scientists who received training through the IGGI, including Paul Mireji, would later apply for their own grants to lead new workshops.
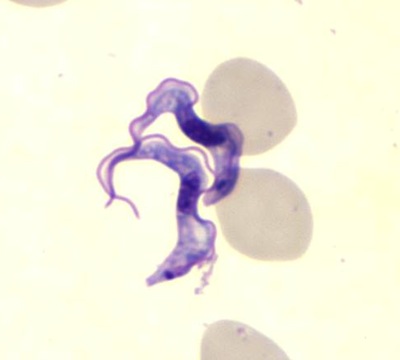
A blood stain reveals two specimens of Trypanosoma brucei. Image credit: Blaine Mathison, Centers for Disease Control and Prevention
Bioinformatics is a strength of scientific institutes in countries like Kenya and South Africa, where computational infrastructure is more available than the expensive and difficult-to-maintain equipment used for many wet lab experiments. Nonetheless, as raw DNA data about the tsetse fly started to come in, African centers often had to improvise solutions to problems that rarely occur in Europe or the U.S.
“We don’t have the kind of bandwidth that you get in the States,” says Mireji. “So we’ve been able to develop our own local, standalone tools that can work without the use of the Internet.” Bioinformaticians typically make high demands of the Internet, but to help assemble the G. morsitans genome and make comparisons with other species, Mireji and his colleagues had to put together a fully offline cluster with specific tools already installed. They also needed their own, local copies of the DNA data they were using, including both the fragmentary G. morsitans genome and the completed genomes of other insect species. (The BRI has since resolved most of its connectivity problems.)
In the end, the model for the project was one of pooled resources and shared data, where wealthy centers abroad produced the genome, and newly trained bioinformaticians across Africa helped make sense of it. Tsetse DNA made its way to the Wellcome Trust Sanger Institute in the U.K., which sequenced the genome piece by piece, often at its own expense. In the U.S., VectorBase, a unit of the National Institutes of Health, stored the data and made it available to IGGI members. But when the G. morsitans genome was finally presented in April 2014, it was accompanied by a collection of 12 scientific papers in the open access journal PLOS Neglected Tropical Diseases, where many of the IGGI’s trainees revealed their discoveries about how the fly’s genome operates.
From assembling a whole genome out of DNA fragments, to investigating the genetics behind behaviors like prey-seeking, milk production, and the transmission of trypanosomes through saliva, the accomplishments of the IGGI were all at least partly centered on the same institutes working to control the spread of tsetse flies in their home countries. “I can’t imagine where funding would come from if I said today, I just want to build capacity in bioinformatics in Africa,” says Aksoy. “The fact that the end goal was to discover pathways and genes that would interrupt sleeping sickness transmission… I think it was instrumental.”
Local Capacity
Although it’s accomplished its headline goal, the IGGI is not winding down. With help from the Genome Institute at Washington University of St. Louis, the project has gone on to produce five more Glossina genomes of different species and subspecies. “We want to understand a little more about the genetics of the tsetse fly in terms of populations, and the gene flow between flies,” says Mireji, who has been analyzing these new genomes at the BRI. “Having that information, we could better understand whether we can adopt similar approaches for control of tsetse flies in different habitats and different regions.”
The IGGI is now part of the weft of genomics in Africa, an increasingly sophisticated enterprise. Collaborations formed through the initiative are still active; Mireji, for instance, has a part-time appointment at Yale, where he studies interactions between flies and trypanosomes. But he views his work in Kenya as a far more relevant model for how tsetse flies will behave in the wild, as his colleagues in the BRI create and maintain modified strains of flies to test their behavior.
“When it comes to functional genomics, that can only be done here,” Mireji says. “In the U.S. and U.K., the tsetse fly samples come from lab colonies. Going through those bottleneck selection processes in the lab, we’re not sure anymore if we’re dealing with a tsetse fly anything like what we have in the field.”
._Coloured_drawing_by_A.J.E_Wellcome_V0022550.jpg)
Illustrator Amedeo John Engel Terzi drew this picture of a tsetse fly in the early 20th century. Our understanding of the fly's biology has expanded enormously in the past 100 years, but the trypanosomiasis it spreads is still a critical public health threat. Image credit: Wellcome Library, London
Even as scientists like Mireji get closer to the genetic origins of crucial traits like immunity and navigation, training the next generation of researchers remains a high priority. With a grant from the U.S. National Institutes of Health, Mireji directs a workshop in bioinformatics that continues to take on new tools and applications. Today, his trainees learn how to assemble genomes from scratch; how to search across genomes of many different flies for genetic variants that might change their behavior; and techniques for dealing with RNA data, which gives a more momentary snapshot of what the fly is doing on a cellular level.
Thanks to swift advances in sequencing technology, scientists in several African countries are increasingly able not only to interpret genetic information, but also to produce it. While the work of sequencing entire genomes is mainly farmed out to the large international centers, several institutes in South Africa, Kenya, Uganda, and elsewhere have acquired their own sequencers for RNA studies. The BRI’s sequencers come from an unorthodox source called Seeding Labs, which distributes scientific equipment to build local capacity, rather than tying its programs to specific projects.
“The good thing about Seeding Labs is that this equipment is not just for one institution,” says Mireji. “We are sharing the equipment, so the equipment based at Gulu University [in Uganda], and at Egerton University and Nairobi University here, is infrastructure we are sharing.”
That kind of regular, cross-institutional collaboration is a hallmark of modern genomics, which deals with volumes of data, and webs of analysis, too large and complex to keep in-house. In the fight to control trypanosomiasis, data is likely to cross many borders, as a DNA sequence becomes a genetic variant, then an engineered strain of flies, and finally, with luck, a new intervention — one that keeps flies away, causes their development to fail, or foils the ability of T. brucei to establish itself in its tenacious hosts.
The G. morsitans genome produced by the IGGI is only the first and simplest step in that process. But the international structures created for the project, and the young scientists who got their first experience working with DNA data through its workshops, are carrying through.
“My goal would be for there to be more of these efforts,” Aksoy concludes. “I still think Africa is a great place to build this kind of capacity. There is less dependence on wet lab kinds of resources, and they’re very good people. They’re good centers, and there’s close communication between the computational biologists and the wet lab scientists.”


