All in One Day by 2020
May 11, 2016 | Contributed Commentary | Ketan Paranjape general manager of the Health & Life Sciences Group, Intel Corporation, welcomed the crowds at the Bio-IT World Conference & Expo last month in Boston. Paranjape’s opening address was a call to action for the Bio-IT community: harness big data and technology to achieve new heights for precision medicine. Here, Ketan lays out Intel’s goals for precision medicine and how the Bio-IT community can join in. —The Editor
Today, precision medicine is typically available only to select patients at sophisticated oncology centers—and with farsighted governments or generous insurance policies to cover the costs. The pipeline of steps from initial diagnosis to solid treatment plans often takes weeks or months to move through, leaving patients and their families mired in anxiety and uncertainty. And the universe of known targeted therapeutics is still woefully small.
At Intel, we believe that by 2020, clinical teams can be empowered to diagnose cancer or another genetically-based disease and conduct secondary analysis that identifies causal genes and resulting pathways to disease expression—all within 24 hours. We’re calling this vision “All in One Day”, and we’re working with health and life science leaders to make it a reality.
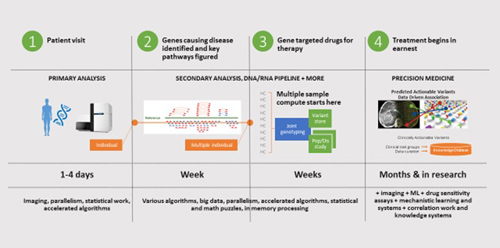
Capturing the Opportunities of Precision Health Analytics
Nations around the world are engaged in genomics-based programs to improve population health. In addition to the United States’ $1 billion National Cancer Moonshot, the National Institutes of Health is enrolling one million volunteers to serve as a research cohort. The People’s Republic China is finalizing plans for a large-scale genome sequencing program focusing on specific cancers. Genomics England is sequencing the genomes of 100,000 National Health Service patients who suffer from rare genetic disorders. And the list goes on.
The biggest technology roadblocks to All in One Day arise from the need to manage, pool, analyze, and store the enormous quantities of genomics and other data.
The human genome itself is 3 billion base pairs, and the raw sequencing data is just a start. The output from genome sequencers must be assembled, analyzed, compared, and studied. With the number of sequencers rising, their performance climbing, and costs falling, a study published in PLoS Biology (doi:10.1371/journal.pbio.1002195) predicts that by 2025, the data requirements for genomics alone will equal or surpass those of the entire field of astronomy or all the content on YouTube.
Data diversity adds to the challenges. Before clinicians can formulate practical treatment plans, researchers must combine genomic data information from privacy-protected electronic health records as well as a wide variety of clinical, pharmaceutical, epidemiological, and other data. As the Internet of Things gathers steam, researchers will incorporate data from billions of lab instruments, molecular imaging systems, clinical monitoring platforms, implantable diagnostics devices, personal health devices, and other sensor-based and embedded technologies.
Beyond the volume and variety of data, precision medicine researchers must remove institutional and technical barriers to secure data analysis. Data is often siloed in discrete databases at multiple organizations, making it difficult or impossible to pool data sets and perform the comprehensive, collaborative analysis that leads to lifesaving clinical breakthroughs.
To build success, we must empower researchers and clinicians to securely share data across institutions while protecting patient privacy. Computing architectures and technologies, as well as tools and applications, must advance to support the rising requirements and diverse new workloads that will be part of the precision medicine ecosystem. Interoperable solutions must be based on open industry standards to foster collaboration.
Tools and Collaborations to Address the Challenges
As an innovation catalyst for the technology industry, Intel is collaborating with life science and technology leaders to solve these challenges. In addition to advancing our own technologies, we’re supporting the development of collaborative ways to share data while keeping it federated and secure. Our engineering teams have worked with open-source developers since 2011, producing major performance gains in widely used life sciences tools, algorithms, and applications.
For example, Intel has worked closely with the Broad Institute of MIT and Harvard to co-develop new tools and capabilities for large genomic workflows. One such innovation is Cromwell, a new workflow engine that simplifies the job of orchestrating genomic workflows to run at cloud scale. Cromwell enables researchers to more easily prepare and run their genomic workflows in a variety of cloud environments, whether those are on-premises private clouds or secure public cloud services. Cromwell uses its built-in intelligence to improve performance by finding optimal ways to execute tasks and the most appropriate hardware resources on which to run those tasks. Broad is also teaming up with Intel, Cloudera, and four leading cloud service providers to enable cloud-based access to its popular Genome Analysis Toolkit* (GATK*) software.
In 2015, Intel and Oregon Health & Science University (OHSU) established the Collaborative Cancer Cloud, an open-source-based analytics platform for precision medicine. This year, Intel and OHSU’s Knight Cancer Center welcomed the Dana-Farber Cancer Institute and the Ontario Institute for Cancer Research as recent additions to the Cancer Cloud. The Collaborative Cancer Cloud enables medical institutions to securely share insights from their private patient genomic data for potentially lifesaving discoveries. The Cancer Cloud’s unique, federated approach to data sharing allows for rapid advances while overcoming many concerns about sharing sensitive datasets. In doing so, it speeds genomic sequencing and enhances clinician productivity, giving physicians more time to work with patients and devise effective, personalized treatments.
Intel is also conducting ethnographic research around the world to understand the attitudes and desires of patients, clinicians, and other stakeholders on key topics related to precision medicine. This research helps us design and inspire solutions that meet the deeply felt needs of patients and their loved ones while also addressing clinician concerns and requirements.
Moving Forward Together
On the technology front, Intel is pursuing dramatic advances to provide performance, power efficiency, throughput, and reliability for the foreseeable future. The umbrella for our advances is something we call Intel Scalable System Framework. This holistic design philosophy aims to meet the computational needs of precision medicine and other fields that depend on high-performance computing to achieve their objectives. The framework’s flexibility helps reduce costs and complexity by allowing systems to run both compute-intensive and data-intensive workloads, as well as machine learning and visualization applications.
All of us—as individuals, governments, policy makers, researchers, healthcare providers, payers, and technology innovators—have a stake in the success of All in One Day. We all have roles to play to build that success. Here are four important actions to accelerate progress:
- Get educated. Learn about precision medicine’s benefits, as well the issues it raises for ethical standards, patient privacy, informed consent, and other matters. Recognize the need to advance big data analytics and secure information sharing as key enablers for precision medicine.
- Get involved. Make your voice heard. Develop and advocate for patient-centered policies that support rapid progress. Collaborate to remove roadblocks.
- Support open standards. Develop and deploy flexible, interoperable technologies and solutions that allow for cost-effective, compatible computing and collaboration from the research center to the oncologist’s office.
- Gather evidence. There’s a compelling business and clinical case for precision medicine, but we need to prove it. In addition to reducing suffering, what are the economic benefits of faster, more precise diagnosis and treatment enabled by whole genome sequencing? How much can we save by getting patients back to their normal lives faster? How will our healthcare system benefit from avoiding unnecessary diagnostic procedures and repeated, trial-and-error treatments?
I look forward to working together to achieve the vision of All in One Day—and move further. All in One Day is a single milestone as we advance toward a transformative era of precision health and wellness and predictive biology.


