Machine Reasoning Extends AI's Scope In Clinical Data Quality And Research
Contributed Commentary by Robert Stanley
July 24, 2019 | How can artificial intelligence (AI) foster accurate, research-quality electronic medical records and complementary study data to provide semantic interoperability across standards? How can this technology create more meaningful data connections that support research and analysis? These are important questions, as electronic medical records (EMR) systems aren’t living up to their promise for creating useful clinical data. EMR data quality problems are creating risks for missed diagnoses, prescriptions, billing. Clinicians are faced with tedious data entry that can benefit by unobtrusive input verification and auto-completion. Coupling traditional data quality methods with AI makes it possible to efficiently solve issues like ensuring correct data, reporting to related systems, identifying medicine interactions, improving care processes—without changing clinical workflows.
Old vs. New
There are several traditional methods for improving data quality in drug discovery: rules-based data quality assessment and transformation; drug terminology normalization to standardized lists or “lexicons” via scored string matching; and statistical analyses of data quality. Combined with innovative technology, these methods themselves become more effective. Traditional lexicons, rules, and statistical analyses can be applied to complement and train AI algorithms, enabling greater efficiency in solving drug development problems. When applied to focused and well-defined applications, the result is a smarter kind of data tool.
As examples, standard AI methods include machine reasoning (MR) and machine learning (ML). MR builds on newer AI-enabling database technologies (i.e., NoSQL, Graph and Semantic) for advanced systems. MR applies “ontologies” which consist of semantically-meaningful data models that can be used for deductive reasoning, entailment, and decision support, even when datasets are incomplete (see, “Filling Gaps”). Ontology-enabled MR can augment lexicon-based text matching by taking context into consideration. If a term is ontologically positioned as a gene (e.g., comprised of amino acids, expresses proteins), the system will know to restrict lexical matching to potential genes, and to flag strings that aren’t identifiable as genes.
AI In The Toolkit
It’s important to remain focused on a specific project when adding AI into the technology mix. Taking on large-scale AI applications may be overly ambitious—and therefore present a roadblock to effective use of the technology. Instead, treat AI as one more option within a broad data quality toolkit.
Using this approach, Parkinson’s Institute and Clinical Center (PICC) transformed data such as unstructured text, XML, tables, tsv, and other data formats into a research quality, well-managed data resource. This helped the organization meet its technical goals including creating a new, unified “Parkinson’s Insight” data resource. Business goals were achieved as well, including researching and publishing discoveries from that data, and engaging in revenue-generating partnerships based on the new data resource.
Blend Modern Technologies In Smart Ways
Overall, data quality poses several challenges in drug research and discovery, development, and delivery—challenges that can be overcome by blending the advantages of the old and the new to resolve common data quality problems.
Differing chemistry terminology can create research and business glitches. Standardized lexicons with synonyms and preferred terms can often be applied to readily transform data from one standard lexicon to another. Knowledge engineers and their advanced data scientist colleagues can access and apply traditional lexicons in tandem with improved semantic ontologies and MR to manage and assimilate internal research with public data. Cleaning and standardizing drug terminology with traditional methods is just step one. Applying MR can further alleviate data quality issues such as:
- Identification—for extracting valuable content from documents or unclean databases
- Assignment—for fixing data integration classes and relationships
- Enrichment—for augmenting gaps with new data
And drug researchers aren’t the only beneficiaries. End users can connect APIs or use web-based lookups that employ AI in the background. This can help them transform inconsistent drug terminology and metadata lists into comprehensive datasets with consistent terminology and rich information.
This near real-time transformation can be extremely helpful. Uploaded “dirty” terms can be automatically mapped to FDA’s preferred terms. Additional data can be added according to interest, such as NDC codes, proprietary and generic names, FDA Product IDs, labeler names, routes, dosages, and associated biological mechanisms, streamlining efforts to achieve clean, rich, usefully connected data.
Maximize Clinical Value By Addressing Practical Challenges
The possibilities are endless—and by focusing on specific research and business bottlenecks, clinicians can most effectively tap into AI’s value. Ideally, AI can be applied to enhance narrowly defined applications including drug terminology normalization, enrichment of existing drug data with published information, and master data integration and management.
These opportunities are likely to bring significant benefits to drug discovery research in terms of reduced time and cost to achieve practical goals. Because of this, AI has the potential to make a positive difference, finally moving drug discovery and development beyond the data quality issues that have been a longstanding hallmark of the life science industry.
Bob Stanley leads Melissa Informatics’ slate of customer projects, blending deep data quality expertise with his robust background as founder of IO Informatics. Bob helps clients harness the entire data lifecycle for business, pharmaceutical and clinical data insight and discovery. Connect with Bob via email at robert.stanley@melissa.com or via LinkedIn.
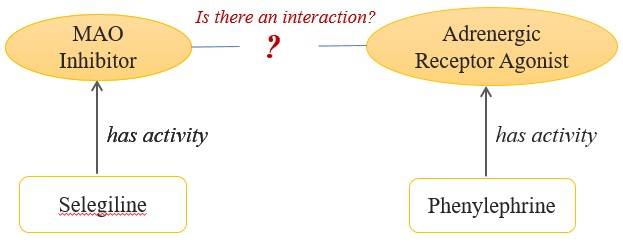
| Filling Gaps How can AI fill in gaps in incomplete datasets? Applying ontologies from machine reasoning creates entailments, or processes that make new assertions about data based on reasoning. Let’s look at a simple example. A researcher’s drug database includes compounds and associated biological activities and mechanisms; however, there is no drug-drug interaction information.
Figure 1: This database can tell researchers that Selegeline is an MAOI and Phenylephrine is an AR agonist but there is no data regarding adverse event risks.
With the database modeled in figure 1, researchers can’t ask if these two drugs are contraindicated. Adding an ontology and MR capabilities solves this significant research challenge.
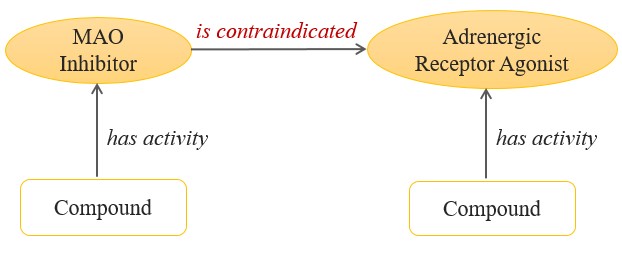
Figure 2: This small ontology, or model of resources and relationships that define an area of interest, asserts that MAOIs are contraindicated with AR agonists. |